Scientists at Harvard Medical School and Boston Children’s Hospital have used a novel gene-editing approach to salvage the hearing of mice with genetic hearing loss, and have succeeded in doing so without any apparent off-target effects as a result of the treatment.
The animals — known as Beethoven mice — were treated for the same genetic mutation that causes progressive hearing loss in humans, culminating in profound deafness by the mid-20s.
The new approach, described online July 3 in Nature Medicine, involves an optimized, more precise version of the classic CRISPR-Cas9 gene-editing system that is better at recognizing the disease-causing mutation seen in Beethoven mice. The refined tool allowed scientists to selectively disable the defective copy of a hearing gene called Tmc1, while sparing the healthy copy.
Notably, the researchers report, their system managed to recognize a single incorrect DNA letter in the defective copy among 3 billion letters in the mouse genome.
Much more work remains to be done before even a highly precise gene-editing therapy like this one could be used in humans, the researchers cautioned. However, they said, the work represents a milestone because it greatly improves the efficacy and safety of standard gene-editing techniques.
“Our results demonstrate that this more-refined, better-targeted version of the now-classic CRISPR/Cas9 editing tool achieves an unprecedented level of identification and accuracy,” said study co-senior investigator David Corey, the Bertarelli Professor of Translational Medical Science in the Blavatnik Institute at Harvard Medical School.
Furthermore, the team said, the results set the stage for using the same precision approach to treat other dominantly inherited genetic diseases that arise from a single defective copy of a gene.
Everyone inherits two copies of the same gene, one from each parent. In many cases, one normal gene is sufficient to ensure normal function that spares the individual from disease. By contrast, in so-called dominantly inherited genetic disorders, one defective copy can cause illness.
“We believe our work opens the door toward a hyper-targeted way to treat an array of genetic disorders that arise from one defective copy of a gene,” said study co-senior investigator Jeffrey Holt, Harvard Medical School professor of otolaryngology and neurology at Boston Children’s Hospital, who is also affiliated with the F.M. Kirby Neurobiology Center at Boston Children’s. “This truly is precision medicine.”
The mice carrying the faulty Tmc1 gene are known as Beethoven mice because the course of their disease mimics the progressive hearing loss experienced by the famed composer. The cause of Ludwig van Beethoven’s deafness, however, remains a matter of speculation.
In mice, the Beethoven defect is marked by one incorrect letter in the DNA sequence of the Tmc1 gene — an A instead of a T — a single error that spells the difference between normal hearing and deafness.
Disabling, or silencing, the mutant copy of the Tmc1 gene would be sufficient to preserve the animal’s hearing, but how could it be done without inadvertently disabling the healthy gene as well?
Two keys are better than one
Classic CRISPR-Cas9 gene editing systems work by using a guiding molecule — gRNA — to identify the target mutant DNA sequence. Once the target DNA is pinpointed, the cutting enzyme — Cas9 — snips it.
So far, these gene editors have shown less-than-perfect accuracy. This is because the guide RNA that leads the Cas9 enzyme to the target site and the Cas9 enzyme that cuts the target DNA are not entirely precise, and could end up cutting the wrong DNA.
To circumvent these challenges, researchers adapted a tool originally developed by Keith Joung, HMS professor of pathology, and Ben Kleinstiver, HMS assistant professor of pathology, both at Massachusetts General Hospital. The adapted tool uses a modified Cas9 enzyme derived from Staphylococcus aureus instead of the standard Cas9 that is derived from the bacterium Streptococcus pyogenes.
To achieve enhanced accuracy of the detection and disruption, the new, optimized system combines two levels of recognition — gRNA to locate the target gene, and a modified form of Cas9 that can pinpoint the specific DNA mutation in Beethoven mice. Using two forms of identification ensures precise and selective cutting of the aberrant copy — and only the aberrant copy — of that gene.
“We took advantage of the fact that this system recognizes mutant DNA but not normal DNA and uses a dual recognition system for enhanced precision,” said study first author Bence György, who conducted the work while at Harvard Medical School and is now at the Institute of Molecular and Clinical Ophthalmology in Basel, Switzerland. “This approach resulted in an unprecedented level of specificity in targeting the mutant gene.”
In an initial set of experiments in cells with and without the Beethoven mutation, the tool accurately distinguished between mutant DNA and normal DNA in copies of the Tmc1 gene. Further analysis revealed that in Beethoven cells, which contained one defective and one normal copy of the gene, at least 99 percent of the molecular “cuts” occurred exclusively in the defective copy of the gene.
Next, researchers injected the gene-editing treatment into the inner ears of mice with and without the Beethoven mutation. DNA analysis showed that editing activity occurred only in the inner ear cells of mice with the Beethoven defect. No editing changes were detected in cells from the inner ears of treated mice that didn’t have the mutation — a finding that confirmed the tool’s precision.
To determine whether the gene-editing therapy interfered with normal gene function, the scientists stimulated hearing cells — called hair cells — from the inner ears of treated mice that didn’t carry the Beethoven defect. The cells showed unchanged, normal hearing responses, affirming that the gene-editing therapy had no effect on normal gene function.
Silencing Beethoven
To measure whether the therapy worked in animals rather than just in cells, researchers performed the gold-standard test for hearing. They measured the animals’ auditory brainstem responses, which capture how much sound is detected by hair cells in the inner ear and transmitted to the brain.
Without treatment, Beethoven mice typically are completely deaf by 6 months of age. By comparison, mice without the defect retain normal hearing throughout life and can detect sounds at around 30 decibels — a level similar to a whisper.
Two months after receiving the therapy, Beethoven mice exhibited markedly better hearing than untreated siblings carrying the genetic mutation. The treated animals were capable of detecting sounds at about 45 decibels — the level of a normal conversation — or about 16 times quieter than untreated mice. The Beethoven mouse with the greatest hearing preservation was capable of hearing sounds at 25 to 30 decibels, virtually indistinguishable from its healthy peers.
Taken together, the findings demonstrate that the novel gene therapy effectively silenced the defective copy of the gene and salvaged the animals’ hearing from the rapid demise typically seen in the disease.
Because the disease is marked by progressive hearing loss, the researchers assessed the effect of therapy on progression over several months. Researchers administered treatment shortly after birth and tested hearing levels in treated and untreated mice with and without the mutation every four weeks for up to six months. In month one, untreated Beethoven mice could hear low-frequency sounds but had notable hearing loss at high frequencies. By month six after birth, untreated Beethoven mice had lost all their hearing. In contrast, treated Beethoven mice retained near-normal hearing at low frequencies, with some showing near-normal hearing even at high frequencies.
Notably, treated animals that didn’t carry the genetic defect did not experience any hearing loss as a result of the gene therapy — a finding that demonstrated the safety of the procedure and its ability to selectively target the aberrant copy of the gene. Even more encouragingly, a small subset of treated Beethoven mice that were followed for nearly a year retained stable, near-normal hearing.
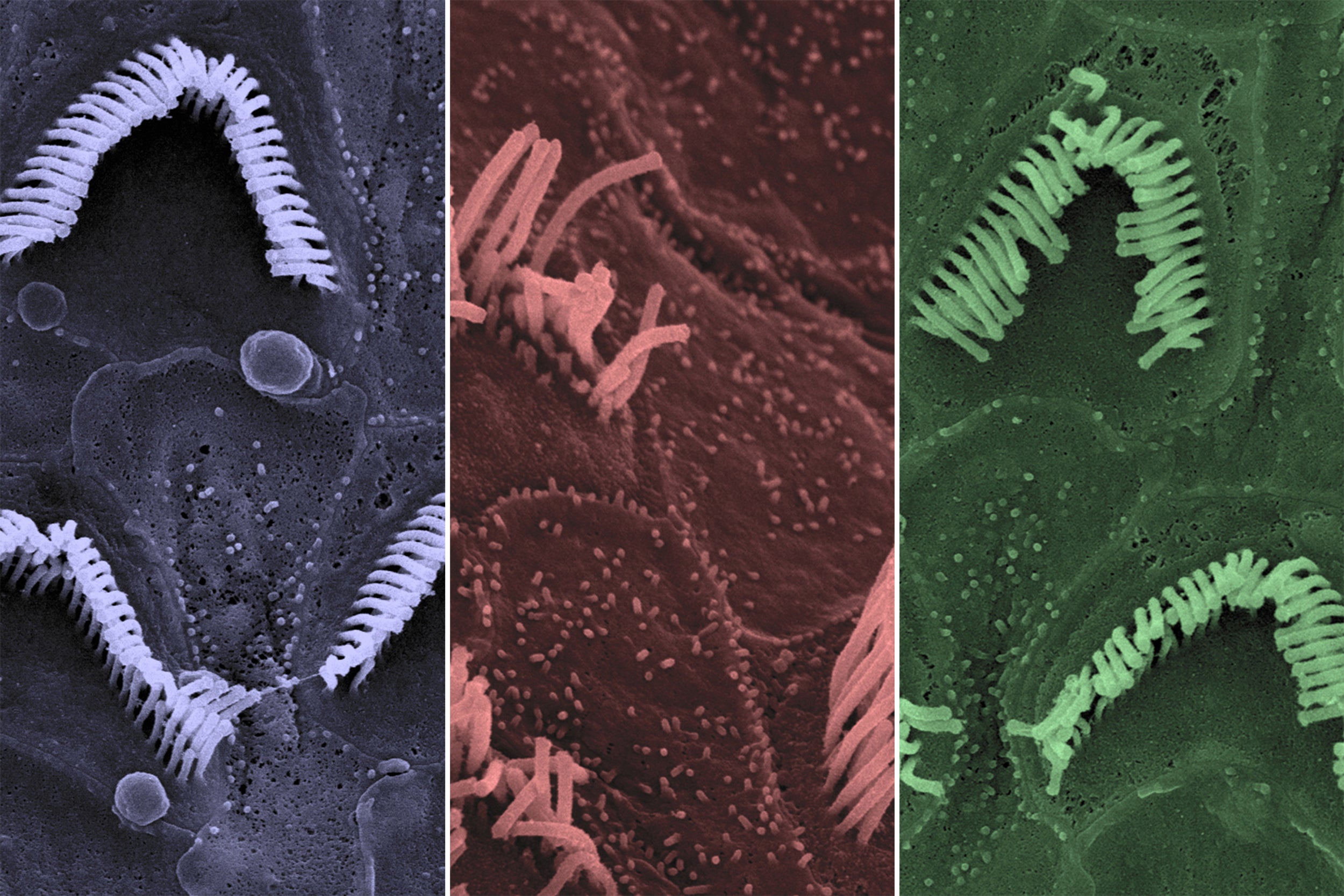
Because the Beethoven defect is marked by progressive deterioration and death of hearing cells in the inner ear, the researchers used electron microscopy to visualize the structure of these critical cells. As expected, in the untreated Beethoven mice, the researchers saw gradual loss of hearing cells along with deterioration in their structure. By contrast, treated Beethoven mice and treated healthy mice both retained a normal number of hearing cells with intact or near-intact structure.
In a final experiment, the scientists tested the effect of the treatment in a line of human cells carrying the Beethoven mutation. DNA analysis revealed that treatment caused editing exclusively in the mutant copy of the Tmc1 gene and spared the normal one.
Because of its ability to target single-point genetic mutations, the approach holds promise for 15 other forms of inherited deafness also caused by a single-letter mutation in the DNA sequence of other hearing genes.
Additionally, the team said, the technique could be adapted for use in other dominantly inherited genetic diseases caused by single-point mutations. To determine its hypothetical utility, the scientists scanned the federal ClinVar database, a national repository of all known genetic mutations linked to human diseases. The analysis showed that based on the tool’s specificity, it could correctly identify 3,759 defective gene variants that are collectively responsible for one-fifth of dominant human genetic mutations.
“To be sure, this is the first step in a long journey,” Holt said. “But what we have here is proof of principle that demonstrates this highly specific, highly targeted treatment could be developed to selectively silence genes that carry single-point mutations and potentially treat many other forms of human disease.”
Co-investigators in addition to Kleinstiver and Joung included Carl Nist-Lund, Bifeng Pan, Yukako Asai, K. Domenica Karavitaki, Sara Garcia, Mikolaj Zaborowski, Paola Solanes, Sofia Spataro, Bernard Schneider, and Gwenaëlle Géléoc.
The work was funded by the Jeff and Kimberly Barber Fund, NIH grants R01-DC013521, R01-DC05439, R01-DC000304, R01-DC016932, DC008853, K99-CA218870, by the Charles A. King Trust, a Banting Postdoctoral Fellowship, the Richard Floor Biorepository Fund, the Kosciuszko Foundation, and the Bertarelli Foundation. Additional support came from a Desmond and Ann Heathwood MGH Research Scholar award and Edward R. and Anne G. Lefler Postdoctoral Fellowship.


